PRinS3 Version 3.0!
Where Innovation Meets Precision!
Revamped Visualization
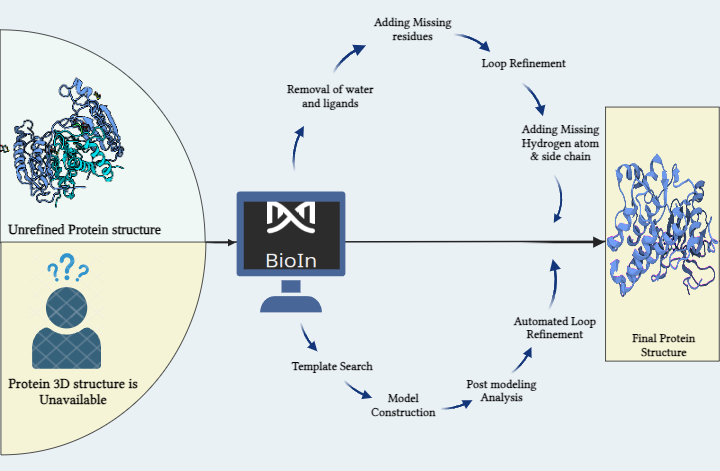
Introducing an upgraded platform designed to enhance the exploration of 3D protein structures based on user-specified PDB IDs. Key features include interactive rotation, zoom, and customization options such as changing protein chain appearances and background colors. Users can also hide water and ligands, select different display styles (cartoon, ribbon, surface), and capture screenshots of their customized views for easy sharing and analysis.SyMoG/AI – A cutting-edge AI platform for drug discovery, now with enhanced features:
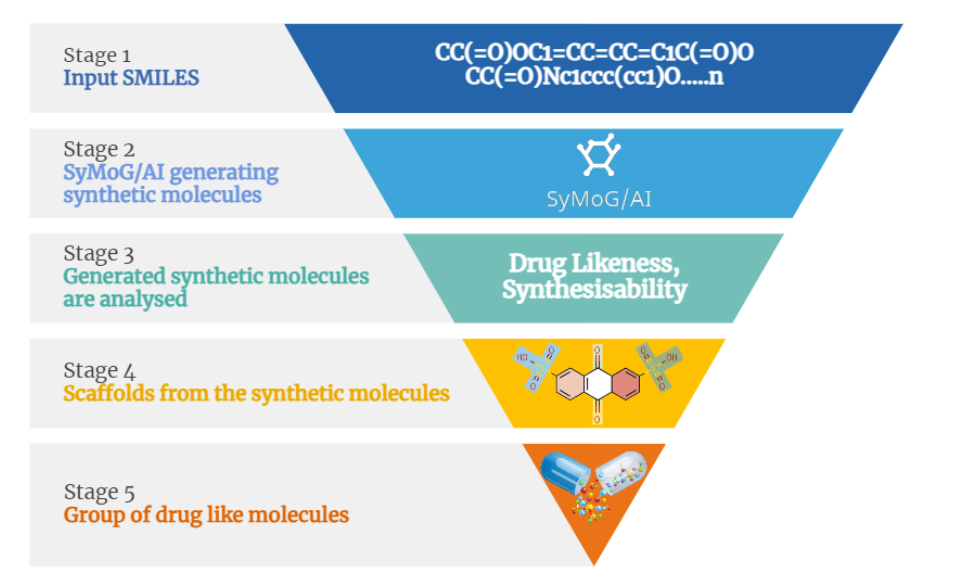
- Target specific is an innovative AI-driven platform that revolutionizes drug discovery by efficiently generating novel synthetic molecules tailored to meet intricate chemical property requirements.
- Property-Based Filtering: Easily filter molecules by specific property values.
- Scaffold-Based Filtering: Filter molecules based on selected scaffolds.
- Export Scaffold Table: Export scaffold data to CSV files.
- 3D Visualization: View and compare multiple molecules in 3D.
- SMILES Format Check: Validate SMILES strings for correct formatting.
X-ESS – A robust platform for molecular analysis with new features:
- Docking Enhancements: Visualize cluster conformations and include metal atoms for accurate metalloprotein simulations.
- MM-PBSA Customization: Manually specify parameters for precise MM-PBSA calculations.
- Molecular Dynamics (MD): Analyze system stability with RMSF, RMSD, and CoM plots, and calculate radius of gyration (Rg). Optimize simulation box size for efficiency.
- pH-Dependent Calculations: Simulate varying protonation states for more accurate results.
Free Energy Surface (FES) Enhancements
The FES module introduces tools to identify the lowest energy conformation, create simulation movies, and download individual simulation trajectories. Users can also customize plot analysis with selective exclusions of specific runs.Optimized for Precision and Innovation in Drug Discovery
With the latest version (PRinS3®), the platform merges cutting-edge AI technology with precise molecular insights to support drug discovery, molecular modeling, and simulation tasks.Request a free confidential discussion and demo today!
PRinS3 streamlines drug discovery with specialized apps. Customize your approach for optimal efficiency and success.
Features
- Explore the deep learning-based generative chemistry app to broaden the chemical search space in drug discovery
- No protein structure? No worries! PRinS3 employs advanced algorithms to generate 3D structures from amino acid sequences.
- Screen multiple compounds against various target proteins simultaneously
- Harness High-Performance Computing seamlessly with PRinS3 - your solution for the cloud or on-premise
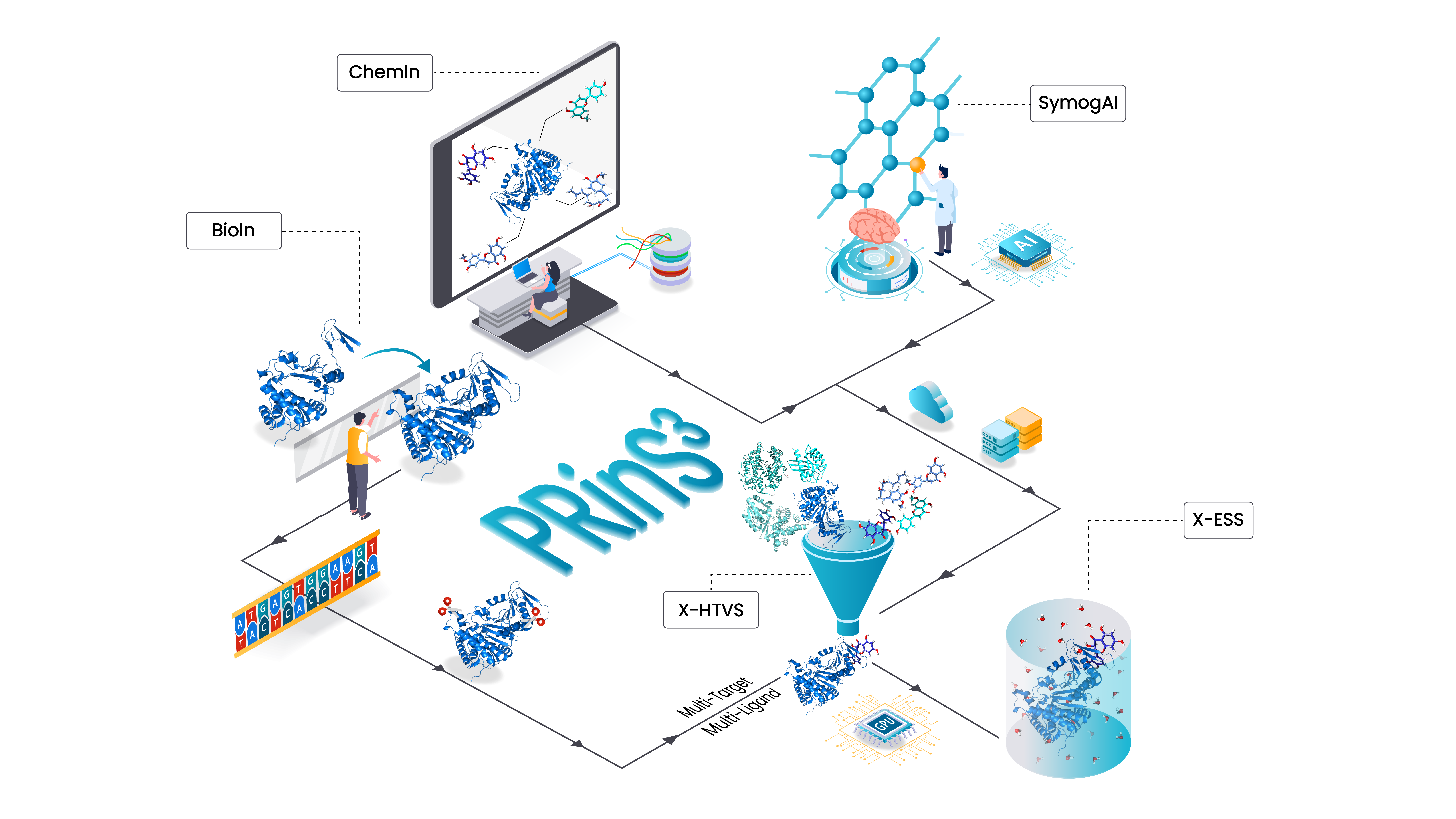
BioIn
Refine and model biological targets fetched from RCSB. Model your protein from sequence and alpha fold database.

ChemIn
Build extensive compound datasets for screening from public databases such as ChEMBL, UniProt, and the natural compound database COCONUT DB. Associate these compounds with targets from UniProtKB/Swiss-Prot, enabling seamless molecule discovery through integrated databases and target-based searches.
SyMoG/AI
Utilize this deep learning-based generative chemistry application to expand the chemical landscape, generating millions of novel drug-like molecules for your research.

X-HTVS
This structure-based application screens multiple compounds against multiple target proteins using a high-throughput virtual process, harnessing the power of cloud computing.

X-ESS
This state-of-the-art application simultaneously screens numerous ligands against multiple protein targets through docking, refining the selection with molecular dynamics simulations and free energy calculations. Our advanced physics-based high-throughput quantitative tool is designed to identify optimal compounds for further synthesis.
QSAR
Ullamco laboris nisi ut aliquip ex ea commodo consequat. Duis aute irure dolor in reprehenderit in voluptate velit esse cillum dolore eu fugiat nulla pariatur. Excepteur sint occaecat cupidatat non proident, sunt in culpa qui officia deserunt mollit anim id est laborum
Lorem ipsum dolor sit amet, consectetur adipiscing elit, sed do eiusmod tempor incididunt ut labore et dolore magna aliqua.
- Ullamco laboris nisi ut aliquip ex ea commodo consequat.
- Duis aute irure dolor in reprehenderit in voluptate velit.
- Ullamco laboris nisi ut aliquip ex ea commodo consequat. Duis aute irure dolor in reprehenderit in voluptate trideta storacalaperda mastiro dolore eu fugiat nulla pariatur.

PRinS3 is a cutting-edge platform harnessing artificial intelligence, physics-based modeling, and cloud computing to streamline and accelerate drug discovery. It provides powerful tools for target identification, molecule design, virtual screening, and property prediction.
PRinS3 delivers solutions tailored for Pharmaceutical Companies & University labs working on drug discovery and computational biology projects.
PRinS3 stands apart due to its:
• AI-driven capabilities: Facilitating faster insights and novel lead generation
• Physics-based foundation: Ensuring predictions grounded in scientific principles
• High-throughput design: Allowing large-scale simulations and rapid screening.
• Focus on customization: Adapting solutions to address industry-specific challenges
PRinS3 is remarkably OS-agnostic, compatible with Linux and Windows. Specific hardware requirements (CPU, GPU, RAM) may vary depending on the computational study but the minimum requirement would be an i5 Intel processor with 8 GB of RAM and 20 GB of storage. For details, please contact Prescience support.
Yes! Please fill out the "Request for Demo" form on the Prescience Insilico website to explore PRinS3's features.
Prescience Insilico takes data security very seriously. PRinS3 employs robust security measures and can be deployed on-premises or on a secure cloud infrastructure based on your preference. Contact us for specifics about our security protocols.
Yes, PRinS3 offers solutions for target identification and validation, helping you explore novel pathways and disease mechanisms.
PRinS3 features AI models trained on large datasets of molecular structures. These models generate new, chemically viable molecules with enhanced potential for desired biological activity.
Prescience offers flexible pricing models to fit your needs. You can choose from license-based pricing or pay-per-use options. Contact sales for personalized pricing information.
Prescience provides dedicated technical support throughout your PRinS3 journey. Our team will assist you with installation, onboarding, and addressing any technical queries.
If you have any further questions, please don't hesitate to contact us at ‘support@prescience.in’.